To apply the theoretical models for brain tissue behavior to relevant clinical problems, we solve the governing equations numerically using the open source finite element library deal.ii. Firstly, we use the computational models to carefully calibrate the model parameters through an inverse parameter identification scheme. This is especially important as, for extremely soft materials such as brain tissue, it is virtually impossible to achieve a homogeneous deformation state during mechanical loading. Once calibrated and validated, the computational tools can be used to tackle scientific issues on the borderline between medicine, biology and mechanics:
The role of mechanics during brain development
The characteristically folded surface morphology is a classical hallmark of the mammalian brain, as illustrated in Figure 1. During development, the initially smooth surface evolves into an elaborately convoluted pattern, which closely correlates with brain function and serves as a clinical indicator for physiological and pathological conditions. Recent evidence confirms that physical forces play an important role in pattern selection. Studying the correlation between microscopic biological processes occurring within the brain during embryonic life and the macroscopic forces generated is essential for understanding the mechanism behind both normal and abnormal cortical folding. Using our computational model, we investigate essential determinants of brain folding, whether they are of geometrical, mechanical, or biological nature, encompassing cortical thickness, brain geometry, local tissue stiffness, cell division, and cell migration (see Figure 1).
Combining physics and biology holds promise to further advance early diagnostics of cortical malformations and to improve treatment of neurodevelopmental disorders such as epilepsy, autism, and schizophrenia.
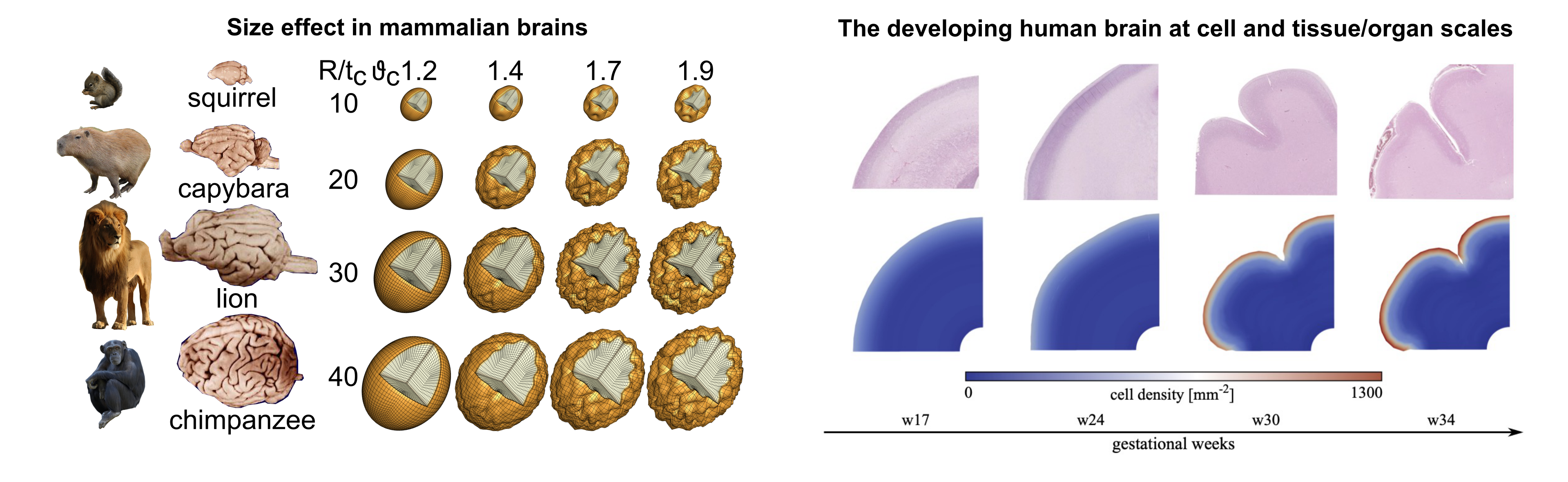
Figure 1: Computational mechanics explain cortical folding patterns in the mammalian brain.
Full scale brain simulations
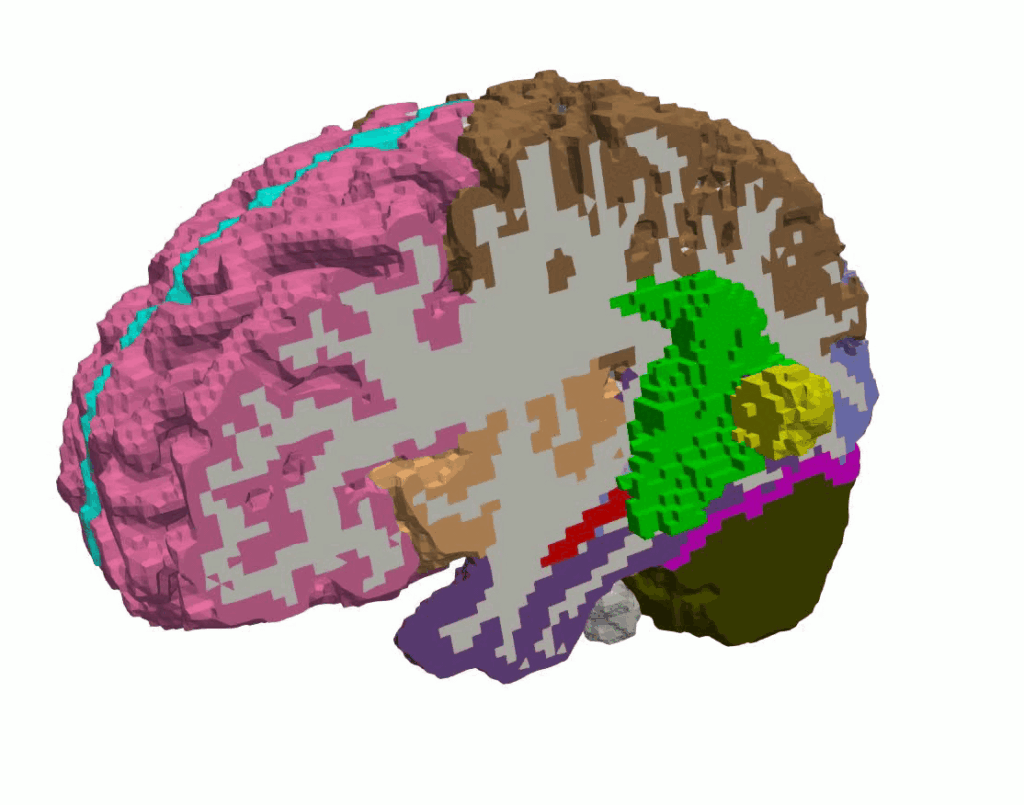
To use the findings from our experimental and modeling investigations in real world applications, we perform simulations using full scale 3-dimensional brain models, as illustrated in Figure 2. These models are created from subject-specific MRI images using the FreeSurfer software suite and an inhouse python code. The resulting models capture the main sulci and gyri of the brain and can be segmented into various regions with distinct mechanical properties.
An important focus of our work is to investigate how different regions with different material properties affect the response of the model under various loading types. We have performed simulations of various surgical procedures (such as brain retraction) and biological processes (such as cerebral atrophy and tumour growth).
By combining our experimental understanding at the tissue level and applying it to a full scale brain model, we hope to contribute to improving the development of protective gear, and to assist the planning of brain surgeries as well as diagnosis and treatment of neurological disorders.
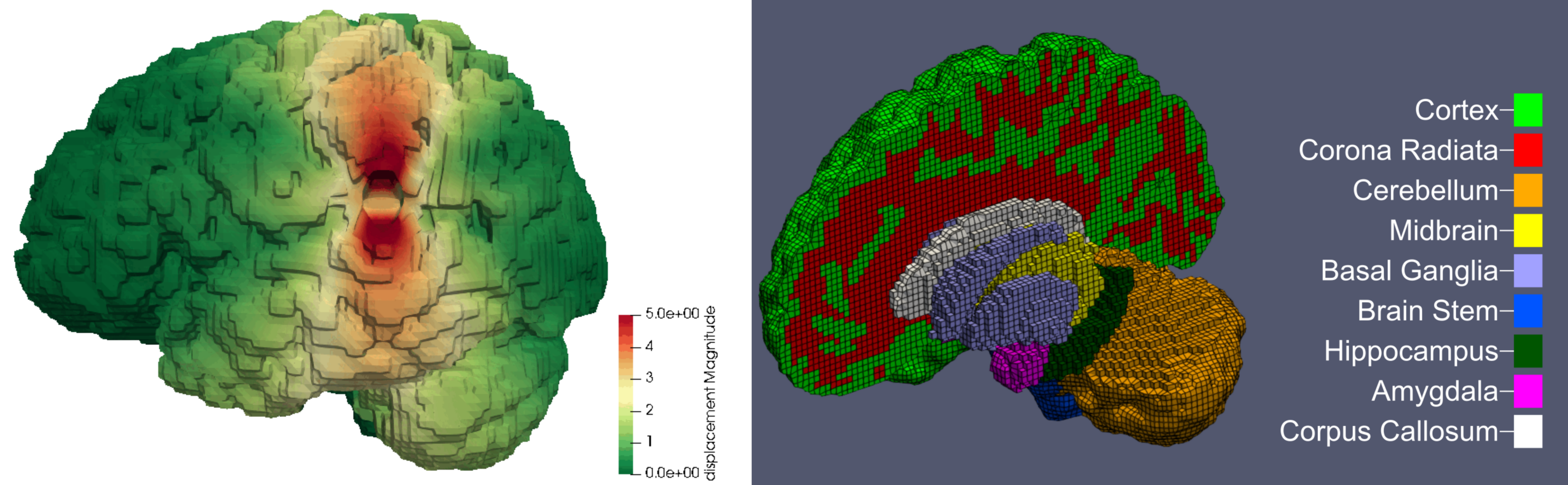
Figure 2: Brain model with regional segmentation
3D Bioprinting
Computational simulations can help predict and tune the mechanical properties of 3D bioprinted constructs – even before fabrication – to save valuable time and cost, as summarized in Figure 3. Like this, a better and faster match between the properties of the biofabricated construct and the target tissue can be achieved. We use the finite element method (FEM) to predict the mechanical properties of different hydrogel mesostructures fabricated through various print patterns and validate our results based on corresponding large-strain experiments under multiple loading modes. This methodology can help save time, material, and cost for future biofabrication applications.
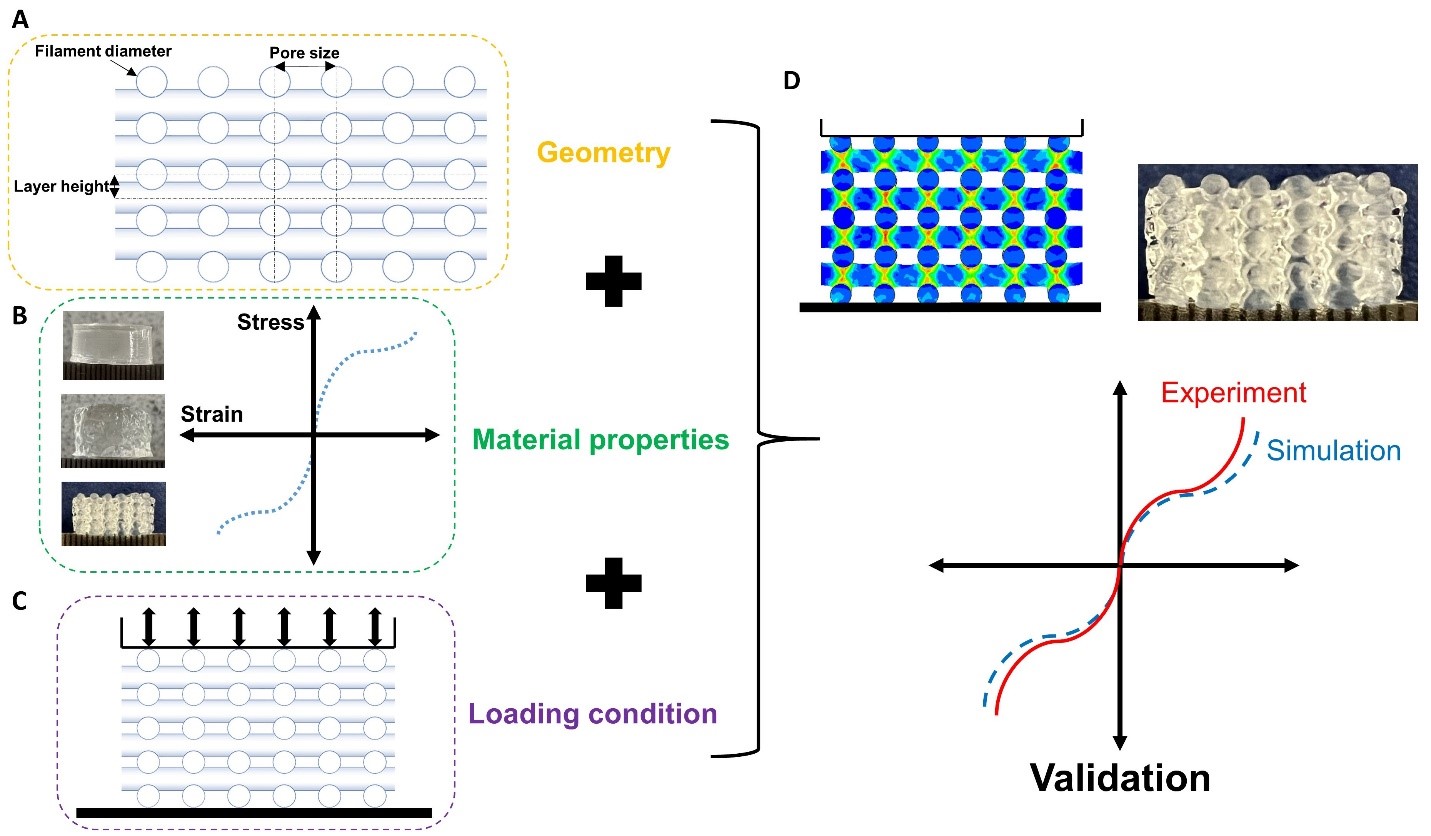
Figure 3: Required steps for developing and validating finite element models to predict the mechanical properties of 3D bioprinted tissue constructs.
Three-dimensional brain models
To translate our experimental and modeling findings to clinically relevant scenarios, we generate full-scale, three-dimensional brain models from segmented MRI data using the FreeSurfer software suite. These anatomically realistic 3D meshes serve as the common computational basis for different applications to simulate brain mechanics under pathological and surgical loading conditions.
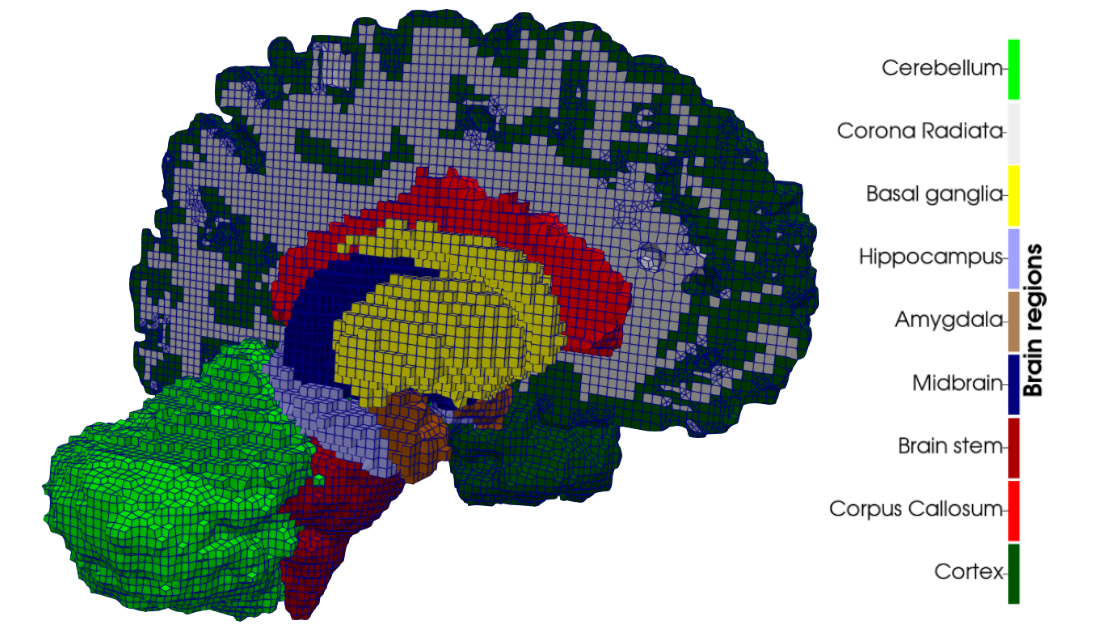
Figure 4: Regionally segmented 3D hexahedral brain mesh generated from MRI data.
Cerebral atrophy
To study the mechanical consequences of cerebral atrophy, we use full-scale brain models in which regional tissue loss is represented through controlled volumetric shrinkage. Based on experimentally informed material parameters, atrophy is modeled as a gradual reduction in volume that mimics neurodegenerative processes such as Alzheimer’s disease. This approach enables analysis of how regional stiffness heterogeneity influences brain deformation, ventricular enlargement, and stress redistribution during progressive tissue loss, supporting interpretation of mechanically driven morphological changes observed in cerebral atrophy.
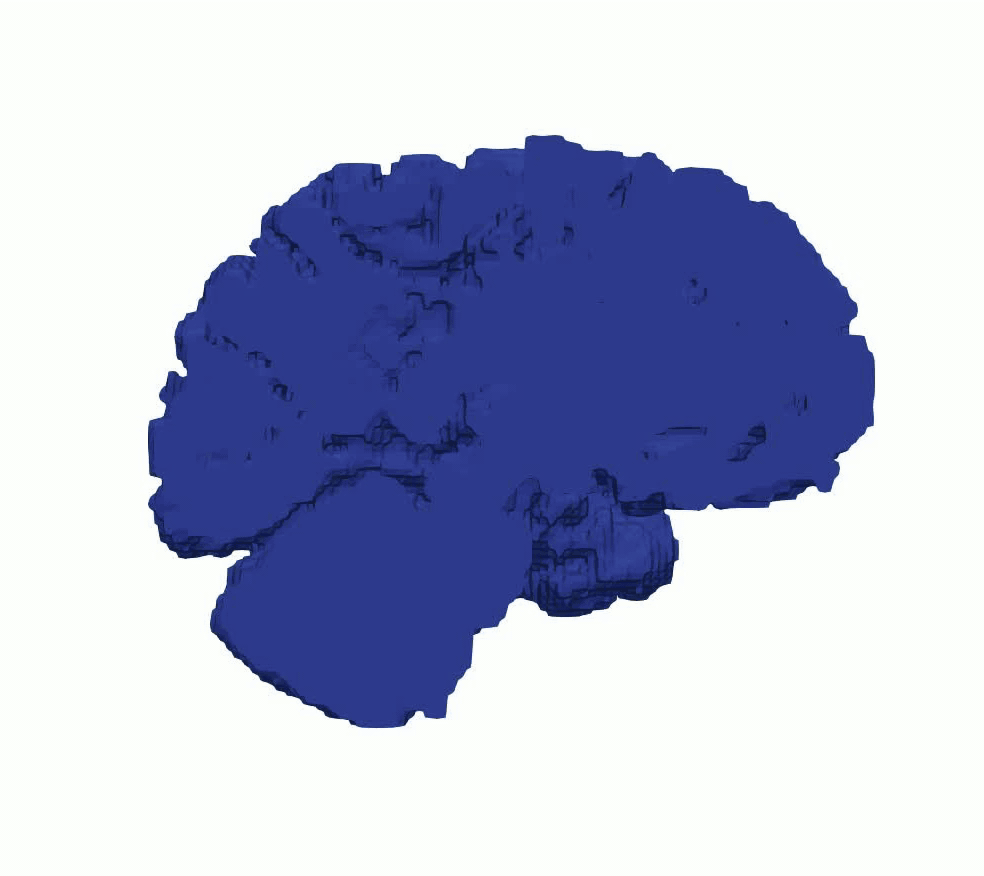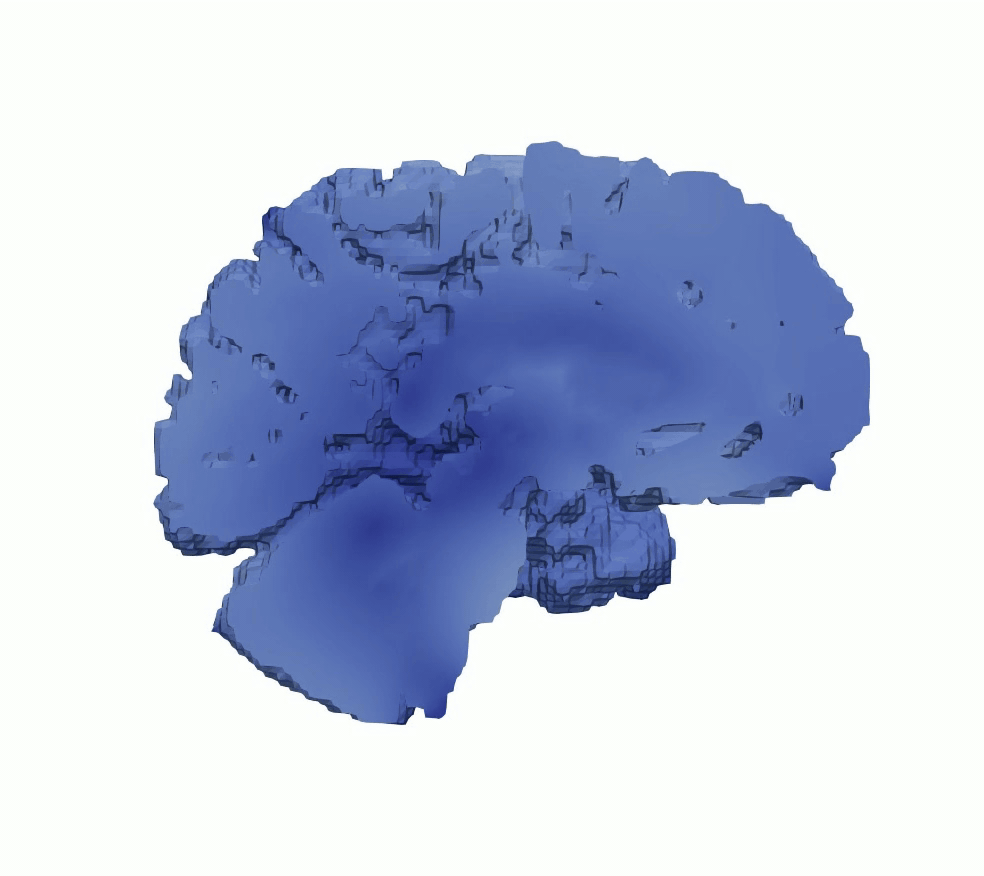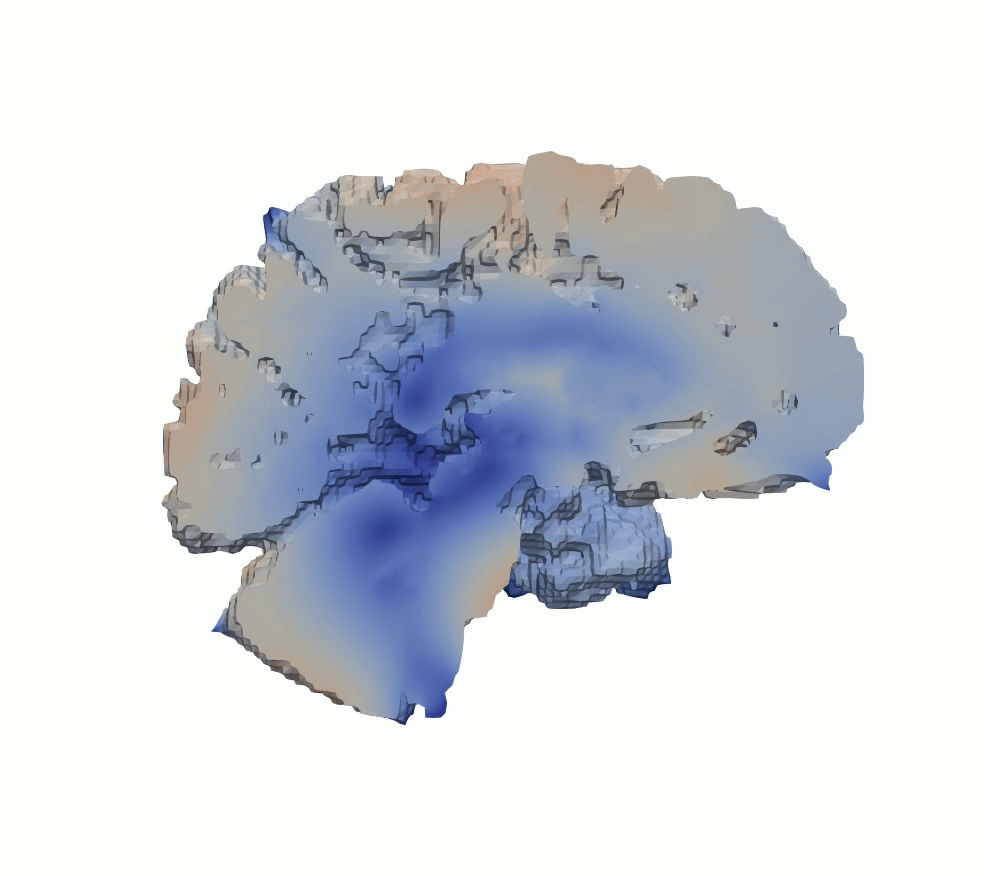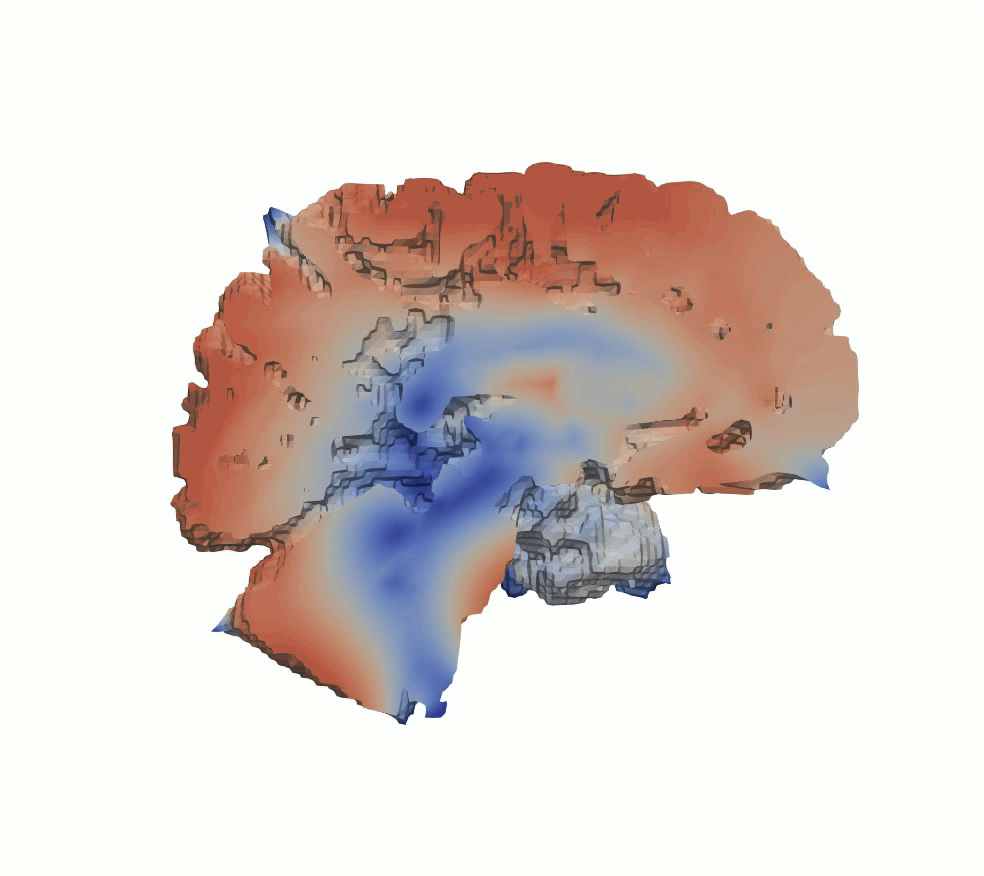
Figure 5: Cerebral atrophy simulated by progressive volumetric shrinkage.
Tumor growth
In the context of tumor growth the investigation of the effect of tumor stiffness, size, shape, and location on brain deformation, stress distributions, and clinically relevant features such as midline shift, ventricular compression, and tumor-induced brain herniation.
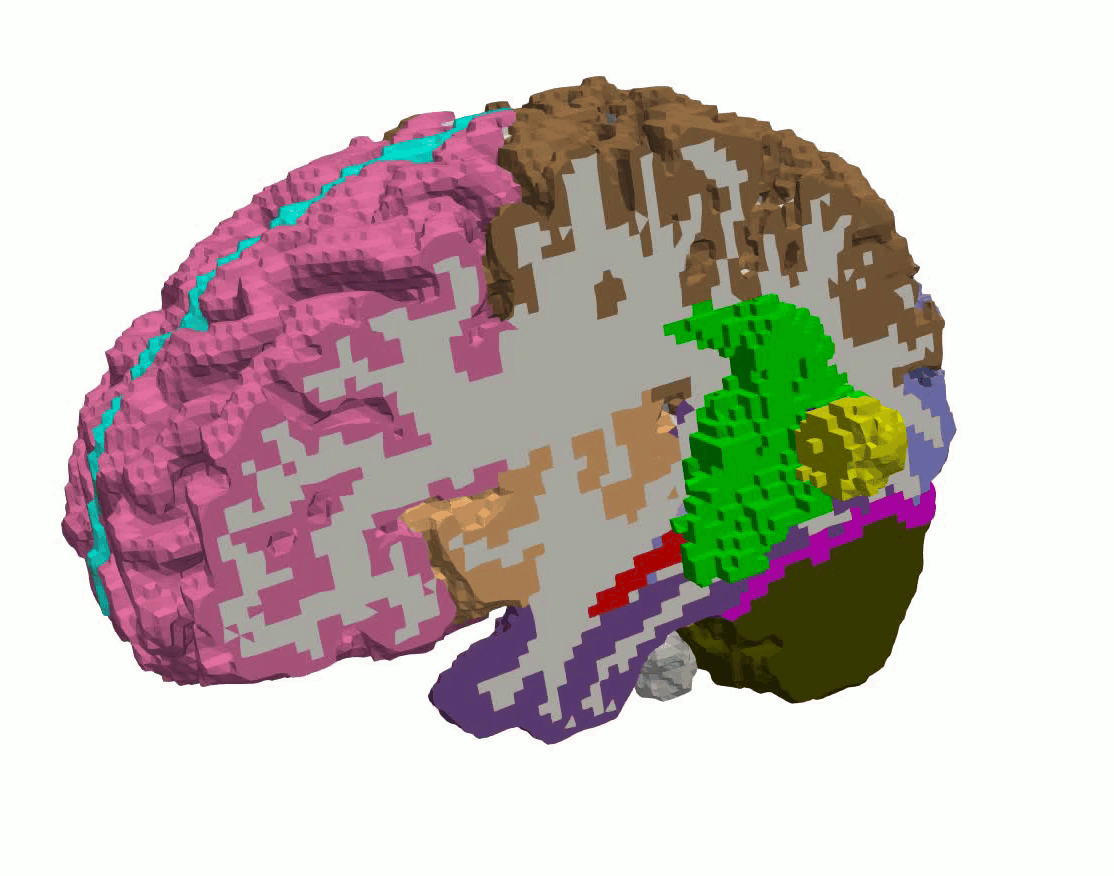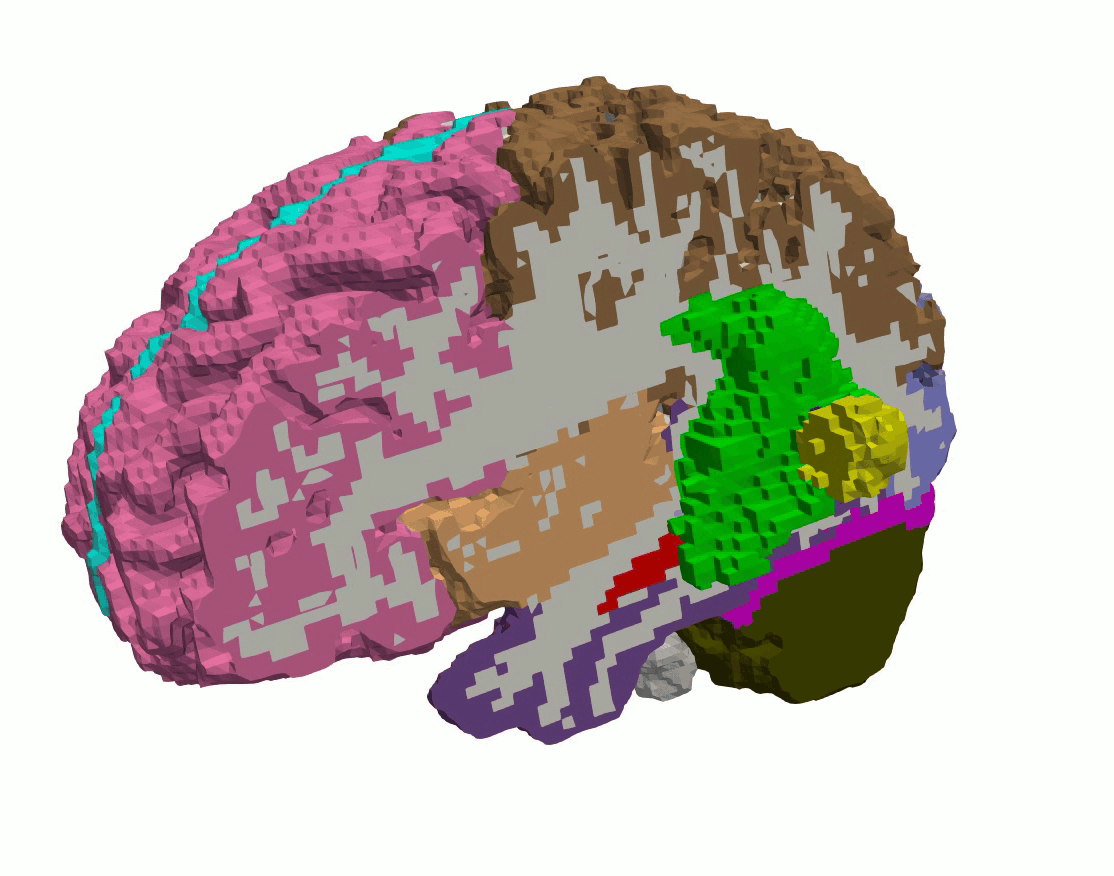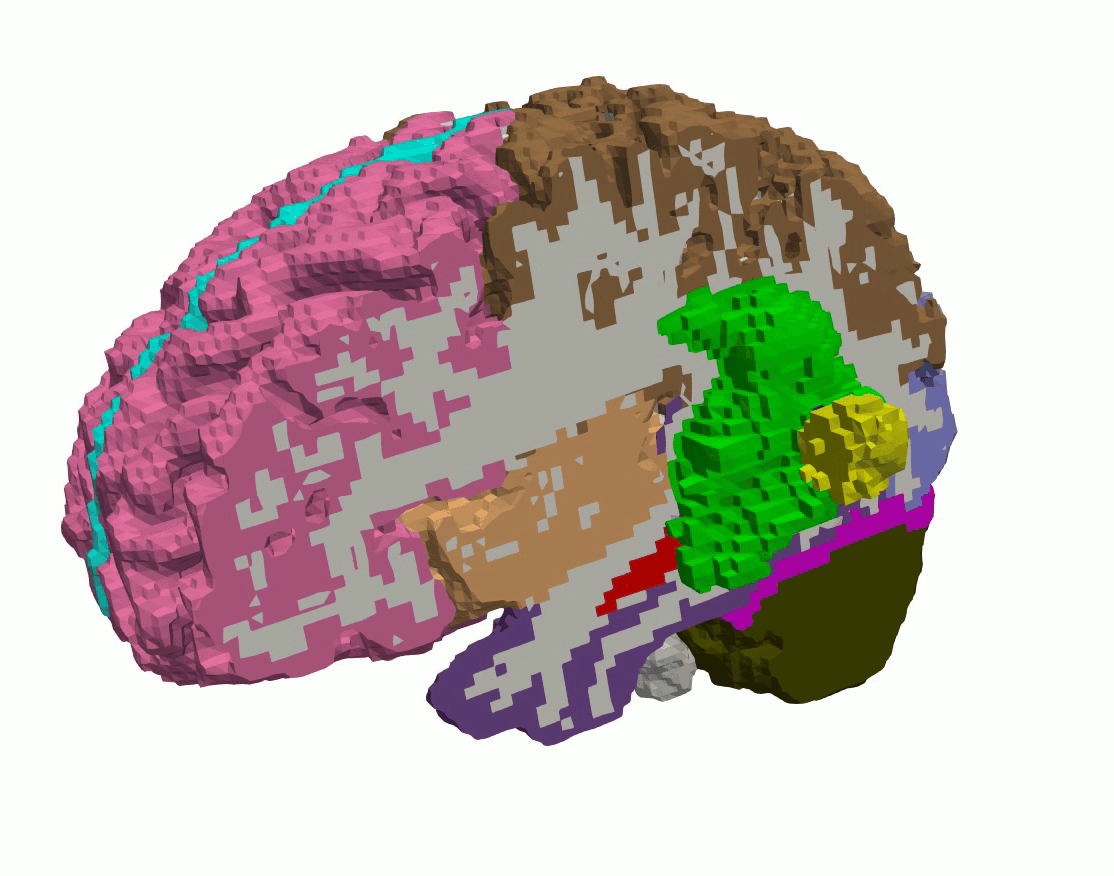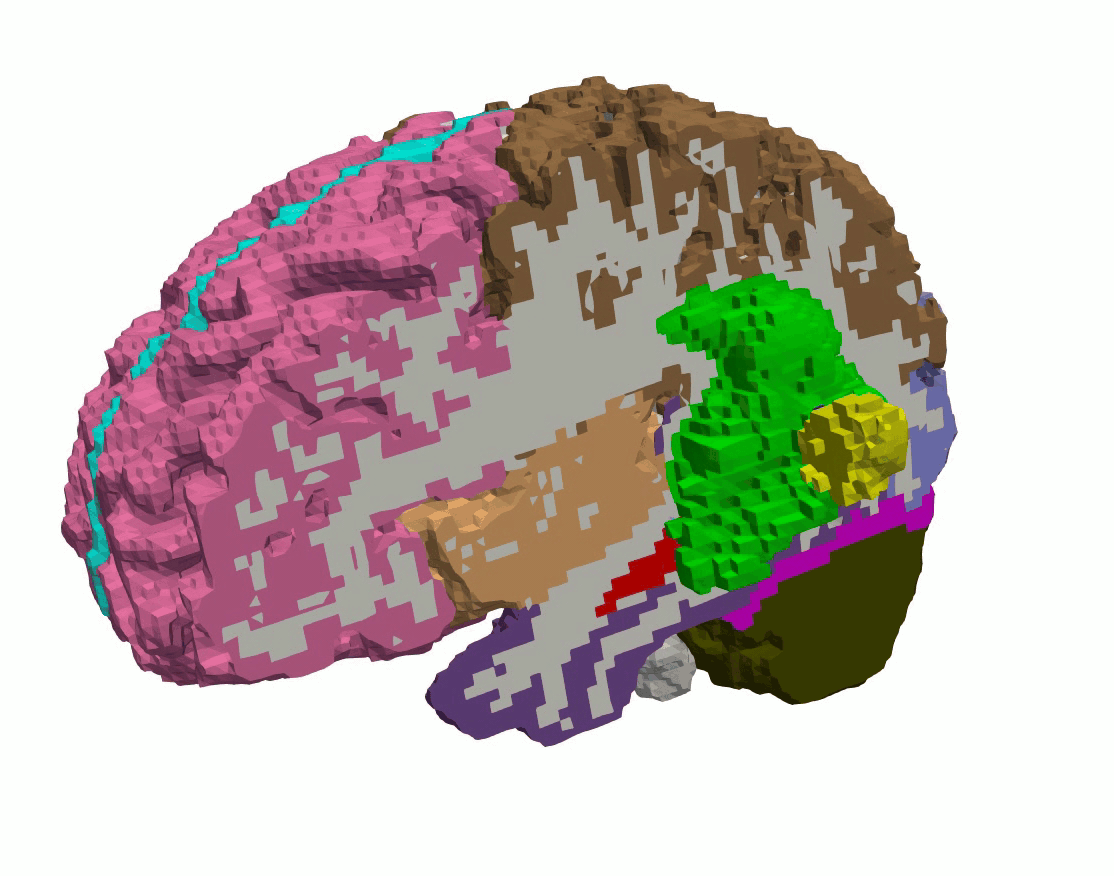
Figure 6: Patient-specific tumor growth simulated by volumetric expansion.
Surgical applications
Beyond pathological processes, the same modeling framework can be applied to simulate neurosurgical procedures, including brain retraction and tissue manipulation during tumor resection. By explicitly accounting for regional material heterogeneity and realistic brain geometry, these simulations enable prediction of tissue deformation, strain localization, and stress accumulation resulting from surgical loading. Ultimately, this work aims to support preoperative planning and the development of safer surgical techniques by identifying mechanical conditions that minimize tissue damage and postoperative complications.

Figure 7: Simulation of brain retraction during neurosurgical manipulation.